Table of contents:
Contents
Converting CBU MRI DICOM data to BIDS format
To start working with your MRI data, you need to convert the raw DICOM format data to NIfTI format and organise them according to the BIDS standard. This tutorial outlines how to do that. If you have any questions, please email Dace. The example scripts described in this tutorial are available on our GitHub repository.
To perform the conversion on your CBU MRI data, you need to be logged into CBU Compute Cluster.
Where is your raw data
The raw data from the CBU MRI scanner are stored at /mridata/cbu/[subject code]_[project code]. You can see all your participant scan directories by typing a command like this in a terminal (replace 'MR09029' with your project code):
ls -d /mridata/cbu/*_MR09029/* /mridata/cbu/CBU090817_MR09029/20090803_083228/ /mridata/cbu/CBU090924_MR09029/20090824_095047/ /mridata/cbu/CBU090928_MR09029/20090824_164906/ /mridata/cbu/CBU090931_MR09029/20090825_095125/ ...
The first part of the folder name which follows the project code, is the data acquisition date. E.g., the data of the first subject in the example above was acquired on Aug-03-2009 ('20090803'). Each subject's folder contains data like this:
ls /mridata/cbu/CBU090817_MR09029/20090803_083228 Series_001_CBU_Localiser/ Series_002_CBU_MPRAGE/ Series_003_CBU_DWEPI_BOLD210/ Series_004_CBU_DWEPI_BOLD210/ Series_005_CBU_DWEPI_BOLD210/ Series_006_CBU_DWEPI_BOLD210/ Series_007_CBU_DWEPI_BOLD210/ Series_008_CBU_DWEPI_BOLD210/ Series_009_CBU_DWEPI_BOLD210/ Series_010_CBU_DWEPI_BOLD210/ Series_011_CBU_DWEPI_BOLD210/ Series_012_CBU_FieldMapping/ Series_013_CBU_FieldMapping/
Each Series### folder contains DICOM files of the particular scan. The name of the folder is the same as what a radiographer named the scan in the MRI console. Usually, the name is very indicative of what type of scan it is. Ideally, you'd also know yourself what type of scans were acquired for your project. In the example above, we acquired a T1w anatomical/structural scan (MPRAGE), nine functional scans (BOLD), and two field maps. The 'Series_001_CBU_Localiser' scan is a positional scan for the MRI and can be ignored.
Each of these folders contains DICOM (.dcm) files, typically, one file per slice for structural scans or one file per volume for functional scans. These files contain a header with the metadata (e.g., acquisition parameters) and the actual image itself. DICOM is a standard format for any medical image, not just the brain. To work with the brain images, we need to convert the DICOM files to NIfTI format which is a cross-platform and cross-software standard for brain images. Along with having the files in NIfTI format, we need to name and organise them according to BIDS standard.
Several DICOM-to-BIDS conversion tools exist (see a full list here). We recommend using HeuDiConv.
DICOM to BIDS using HeuDiConv
HeuDiConv is available on our computing system as an Apptainer image:
/imaging/local/software/singularity_images/heudiconv/heudiconv_latest.sif
HeuDiConv converts DICOM (.dcm) files to NIfTI format (.nii or .nii.gz), generates their corresponding metadata files, renames the files and organises them in folders following BIDS specification.
The final result of DICOM Series being converted into BIDS for our example subject above is this:
├── sub-01 │ ├── anat │ │ ├── sub-01_T1w.json │ │ └── sub-01_T1w.nii.gz │ ├── fmap │ │ ├── sub-01_acq-func_magnitude1.json │ │ ├── sub-01_acq-func_magnitude1.nii.gz │ │ ├── sub-01_acq-func_magnitude2.json │ │ ├── sub-01_acq-func_magnitude2.nii.gz │ │ ├── sub-01_acq-func_phasediff.json │ │ └── sub-01_acq-func_phasediff.nii.gz │ ├── func │ │ ├── sub-01_task-facerecognition_run-01_bold.json │ │ ├── sub-01_task-facerecognition_run-01_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-01_events.tsv │ │ ├── sub-01_task-facerecognition_run-02_bold.json │ │ ├── sub-01_task-facerecognition_run-02_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-02_events.tsv │ │ ├── sub-01_task-facerecognition_run-03_bold.json │ │ ├── sub-01_task-facerecognition_run-03_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-03_events.tsv │ │ ├── sub-01_task-facerecognition_run-04_bold.json │ │ ├── sub-01_task-facerecognition_run-04_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-04_events.tsv │ │ ├── sub-01_task-facerecognition_run-05_bold.json │ │ ├── sub-01_task-facerecognition_run-05_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-05_events.tsv │ │ ├── sub-01_task-facerecognition_run-06_bold.json │ │ ├── sub-01_task-facerecognition_run-06_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-06_events.tsv │ │ ├── sub-01_task-facerecognition_run-07_bold.json │ │ ├── sub-01_task-facerecognition_run-07_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-07_events.tsv │ │ ├── sub-01_task-facerecognition_run-08_bold.json │ │ ├── sub-01_task-facerecognition_run-08_bold.nii.gz │ │ ├── sub-01_task-facerecognition_run-08_events.tsv │ │ ├── sub-01_task-facerecognition_run-09_bold.json │ │ ├── sub-01_task-facerecognition_run-09_bold.nii.gz │ │ └── sub-01_task-facerecognition_run-09_events.tsv │ └── sub-01_scans.tsv
All files belonging to this subject are in the sub-01 folder. The structural image is stored in the anat subfolder, field maps in fmap, and functional images in the func subfolders. Each file is accompanied by its .json file that contains the metadata, such as acquisition parameters. For the functional images, in addition to the metadata files, an events file is generated for each functional run. The file names follow the BIDS specification.
HeuDiConv needs information on how to translate your specific DICOMs into BIDS. This information is provided in a heuristic file that the user creates. At the moment, at the CBU we don't use a standardised system for naming our raw scans in the MRI console. Therefore we don't have a standard heuristic (rules) that we could feed to HeuDiConv for any of our projects. You need to create this heuristic file yourself for your specific project. You can use existing examples as a guideline.
To create the heuristic file, you need to know what scans you have, which ones you want to convert (you don't have to convert all scans, only the ones you need for your project), and how to uniquely identify each scan based on its metadata.
As such, converting DICOM data to BIDS using HeuDiConv involves 3 main steps:
- Discovering what DICOM series (scans) there are in your data
- Creating a heuristic file specifying how to translate the DICOMs into BIDS
- Converting the data
Step 1: Discovering your scans
First, you need to know what scans there are and how to uniquely identify them by their metadata. You could look in each scan's DICOM file metadata manually yourself, but that's not very convenient. Instead, you can 'ask' HeuDiConv to do the scan discovery for you. If you run HeuDiConv without NIfTI conversion and heuristic, it will generate a DICOM info table with all scans and their metadata. Like this:
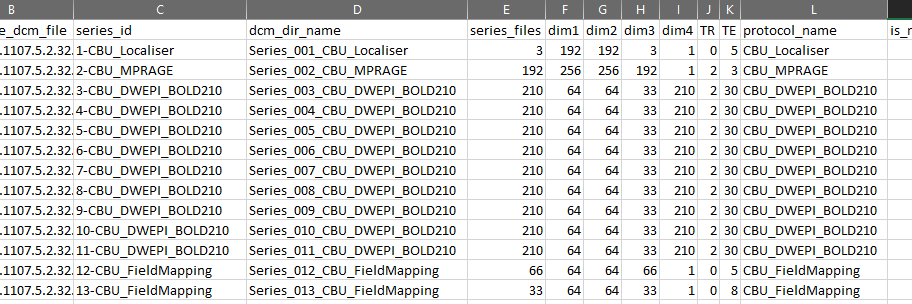
The column names are metadata fields and rows contain their corresponding values.
To get such a table, you'd write a simple bash script, like this dicom_discover.sh script, explained below. You can create and edit the script in any text editor. The file extension for bash scripts is .sh. I named my script dicom_discover.sh.
A bash script must always start with #!/bin/bash.
Specify your project's directory. Below I specified my path for this tutorial.
PROJECT_PATH='/imaging/correia/da05/wiki/BIDS_conversion/MRI'
Specify the raw data directory for any of your project's subjects, let's say the first one. We are assuming here that all subjects in your project have the same type of scans acquired and we can use a single heuristic file for the project.
RAW_PATH='/mridata/cbu/CBU090942_MR09029'
Specify where you want the output data to be saved. I specified the output to go in the 'work' directory. The 'work' directory, in my case, is a temporary directory where I want my intermediate files to go. I will delete this directory once I have finished data preprocessing.
OUTPUT_PATH=$PROJECT_PATH/data/work/dicom_discovery/
Specify this subject's ID. In this case, it can be any, as we will only use this subject to get the information about DICOMs in our project.
subject="01"
To use the HeuDiConv container, you first need to load the 'apptainer' module.
module load apptainer
Now, we can run the apptainer container with the following parameters:
--cleanenv: The HeuDiConv container contains all dependencies internally, it doesn't need any of your environment settings. Therefore, to avoid any potential conflicts with your environment, we specify --cleanenv parameter to ensure that nothing from your environment is passed to the HeuDiConv environment.
--bind: A container is a small, self-contained environment that runs applications. It does not have access to files or folders on your computer. The --bind option in the apptainer command is used to bind/mount files and directories from your system into the container. You must bind everything that the container needs access to. Here we mount our project directory and to the directory where the raw DICOM files are so that the container can access them.
the container image: we specify which container we want to use. In this case the HeuDiConv container located on our imaging system. All parameters after this, are specific to HeuDiConv.
--files: We specify that all our DICOM files are located in the '/mridata/cbu/CBU090942_MR09029/[all subdirectories]/[all subdirectories]/[all files ending with .dcm'
--outdir: Where to output the results.
--heuristic: In this case, we want to discover all scans, not applying any rules.
--subjects: Subject's ID how it will appear in the output.
converter: In this case, we don't want to do the file conversion to NIfTI, we just want to discover what files we have.
--bids: Flag for output into BIDS structure.
--overwrite: Flag to overwrite existing files.
apptainer run --cleanenv \
--bind "${PROJECT_PATH},${RAW_PATH}" \
/imaging/local/software/singularity_images/heudiconv/heudiconv_latest.sif \
--files "${RAW_PATH}"/*/*/*.dcm \
--outdir "$OUTPUT_PATH" \
--heuristic convertall \
--subjects "${subject}" \
--converter none \
--bids \
--overwriteIt's a good practice to unload the module at the end.
module unload apptainer
Once you have created and saved your script, you can execute it. To do that, in a terminal, navigate to the directory where your script is located (I recommend putting your scripts in the [PROJECT_PATH]/code/ directory), e.g., in my case it would be:
cd /imaging/correia/da05/wiki/BIDS_conversion/MRI/code
And the run this command:
./dicom_discover.sh
If everything is working fine, you will see an output like this:
Loading apptainer version:
1.0.3
WARNING: Could not check for version updates: Connection to server could not be made
INFO: Running heudiconv version 0.11.6 latest Unknown
INFO: Analyzing 7145 dicoms
INFO: Generated sequence info for 1 studies with 18 entries total
WARNING: Heuristic is missing an `infotoids` method, assigning empty method and using provided subject id 01. Provide `session` and `locator` fields for best results.
INFO: Study session for StudySessionInfo(locator=None, session=None, subject='01')
INFO: Need to process 1 study sessions
INFO: PROCESSING STARTS: {'subject': '01', 'outdir': '/imaging/correia/da05/wiki/BIDS_conversion/MRI/data/work/dicom_discovery/', 'session': None}
INFO: Processing 18 pre-sorted seqinfo entries
INFO: Populating template files under /imaging/correia/da05/wiki/BIDS_conversion/MRI/data/work/dicom_discovery/
INFO: PROCESSING DONE: {'subject': '01', 'outdir': '/imaging/correia/da05/wiki/BIDS_conversion/MRI/data/work/dicom_discovery/', 'session': None}Once the processing has finished, the table that we are interested in will be located at OUTPUT_PATH/.heudiconv/01/info/dicominfo.tsv. The .heudiconv directory is a hidden directory and you might not be able to see it in your file system. If so, either enable to view hidden files, or copy the dicominfo.tsv to some other location. For example, your home (U: drive) Desktop:
cp /imaging/correia/da05/wiki/BIDS_conversion/MRI/data/work/dicom_discovery/.heudiconv/01/info/dicominfo.tsv ~/Desktop
Now, you can open the file in Excel and keep it open for the next step - creating a heuristic file.
Step 2: Creating a heuristic file
...
Step 3: Converting the data
...
[da05@login-j05 MRI]$ squeue -u da05
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
3246338_0 Main heudicon da05 R 0:06 1 node-k08
3246338_1 Main heudicon da05 R 0:06 1 node-k08
3246338_2 Main heudicon da05 R 0:06 1 node-k08
3246338_3 Main heudicon da05 R 0:06 1 node-k08
More examples
...
Functional scans with `sbref`
...
ls /mridata/cbu/CBU210700_MR21002/20211208_143530 Series001_64_Channel_Localizer/ Series002_CBU_MPRAGE_64chn/ Series003_bold_mbep2d_3mm_MB2_AP_2_SBRef/ Series004_bold_mbep2d_3mm_MB2_AP_2/ Series005_bold_mbep2d_3mm_MB2_AP_2_SBRef/ Series006_bold_mbep2d_3mm_MB2_AP_2/ Series007_bold_mbep2d_3mm_MB2_AP_2_SBRef/ Series008_bold_mbep2d_3mm_MB2_AP_2/ Series009_bold_mbep2d_3mm_MB2_AP_2_SBRef/ Series010_bold_mbep2d_3mm_MB2_AP_2/ Series011_bold_mbep2d_3mm_MB2_PA_2_SBRef/ Series012_bold_mbep2d_3mm_MB2_PA_2/ Series013_fieldmap_gre_3mm_mb/ Series014_fieldmap_gre_3mm_mb/
In this example, researchers acquired a T1w anatomical/structural scan (MPRAGE), five multi-band functional scans with single-band reference scans for each of them, two field maps, and ...
Multiple sessions per subject
...
